|
|
ViewGene |
|
Locus Name:
|
|
|
| GeneID: |
|
|
|
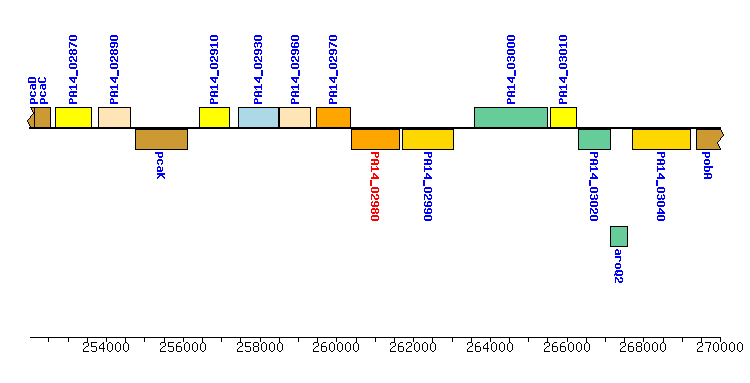
| Location: |
|
|
|
|
|
|
|
|
| Modification Date/Time: | 2006-04-06 13:01:23 |
|
|
|
|
|
|
|
|
|
|
|
|
| Missense Discrepancy: | FALSE |
|
Comments:
Homology: gb|AAG03629.1|AE004462_2 (AE004462) probable porin [Pseudomonas aeruginosa PAO1]
Identities = 419/421 (99%), Positives = 419/421 (99%)
|
Sequence:
ATGTCGCACCCGAGCCACGTCCGCGCGCTCTGCGCTTCCCTCTGCCTGGGCGCCGGCCTGCCAGTCCATGCC
GGGCACGTCCATGCCGGGCAAGGTTTCCTCGAAGATGCCAAGGCCAGCCTCACCGCGCGCAATTTCCACCTG
CACCGCAACTTCGTCGGCGATGCCAGCCAGGGCAAGGCCGAGGAGTGGACGCAGAGCTTCATCCTCGACGCC
CGCTCCGGCTTCACCCAGGGCAGCGTCGGCTTCGGCCTCGATGTCCTCGGTCTCTACAGCCTGAAGCTGGAT
GGCGGCAAGGGAACGGCCGGCACCCAACTGCTGCCAACCCACGACGACGGCCGCCCGGCCGACGACTTCGGA
CGCCTGGCGGTGGCCGGCAAGCTGCGCGTGTCGAACAGCGAACTGAAGATCGGCGAATGGATGCCGGTGCTG
CCGATCCTCCGCTCCGACGATGGCCGCTCGCTGCCGCAGACCTTCCGCGGCGGCCAGCTCAGCGCCAACGAG
ATCGCCGGCCTGACCCTCTACGCCGGCCAGTTCCGCGGCAACAGCCCGCGCAACGACGCAAGCATGCAGGAC
ATGTCGCTGTTCGGCCGGCCAACCGCCACTTCCGACCGCTTCGATTTCGCCGGCGGCGAATATCGCTTCAAT
GGCGAACGCAGCCTGCTCGGCCTGTGGAACGCCGAGCTGAAGGACATCTACCGCCAACAGTACCTGCAACTG
CAACACAGCCAGCCGCTCGGCGACTGGCTGCTGGGCGCCAACCTCGGCGGCTTCCGTGGCCGCGACGCTGGC
AGCGCGCGCGCCGGCAAGCTCGATAACCGCACCGTTTCCGCGCTGTTCTCCGCGCGCTACGGGCTGCATACC
CTCTATCTCGGCCTGCAGAAGGTCAGTGGCGACGATGGCTGGATGCGGGTCAACGGCACCAGCGGCGGAACC
CTGGCCAACGACAGCTACAACGCCAGCTACGACAATCCCGGCGAGCGCTCCTGGCAACTGCGCTACGACTTC
GACTTCGTTGGCCTCGGCCTACCCGGCCTGACCTTCATGACCCGCTACCTGCATGGCGACCATGTGCGCCTG
GCCGGAGTCACCGACGACGGCAGCGAATGGGGACGCGAGAGCGAGCTGGGCTACACGCTGCAGAGCGGCGCC
TTCAAGCGTCTCAACGTCCGCTGGCGCAACTCCAGCCAGCGCCGCGACTGGGGCAGCAATACCCGCTTCGAC
GAGAACCGGCTGATCGTCAGCTATCCGCTGAGCCTGTTGTGA |
Translation:
MSHPSHVRALCASLCLGAGLPVHAGHVHAGQGFLEDAKASLTARNFHLHRNFVGDASQGKAEEWTQSFILDA
RSGFTQGSVGFGLDVLGLYSLKLDGGKGTAGTQLLPTHDDGRPADDFGRLAVAGKLRVSNSELKIGEWMPVL
PILRSDDGRSLPQTFRGGQLSANEIAGLTLYAGQFRGNSPRNDASMQDMSLFGRPTATSDRFDFAGGEYRFN
GERSLLGLWNAELKDIYRQQYLQLQHSQPLGDWLLGANLGGFRGRDAGSARAGKLDNRTVSALFSARYGLHT
LYLGLQKVSGDDGWMRVNGTSGGTLANDSYNASYDNPGERSWQLRYDFDFVGLGLPGLTFMTRYLHGDHVRL
AGVTDDGSEWGRESELGYTLQSGAFKRLNVRWRNSSQRRDWGSNTRFDENRLIVSYPLSLL* |
| AnnotationID:1238 | GeneID:1229 |
|
|
| Modification Date/Time: |
2005-03-23 18:06:37 |
|
|
|
| GeneProduct: | putative outer membrane porin |
|
|
|
| Cell Localization Confidence Code: | 5 |
|
|
| Functional Category: | (15) Membrane proteins |
|
| Alternate Gene Product Name: | |
|
| Functional Category Confidence Code: | 5 |
|
|
| Secondary Functional Category(ies): | transport of small molecules |
|
|
|
|
| Homology: | >gb|AAD39555.1| (AF031417) PhaK-like protein [Pseudomonas putida]
Length = 418
Score = 596 bits (1537), Expect = e-170
Identities = 293/409 (71%), Positives = 341/409 (83%), Gaps = 6/409 (1%)
>gb|AAN67006.1|AE016329_1 (AE016779) BenF-like porin [Pseudomonas putida KT2440]
Length = 418
Score = 595 bits (1534), Expect = e-169
Identities = 292/409 (71%), Positives = 341/409 (83%), Gaps = 6/409 (1%)
>gb|AAO55855.1| (AE016864) outer membrane porin, OprD family [Pseudomonas syringae
pv. tomato str. DC3000]
Length = 418
Score = 569 bits (1467), Expect = e-161
Identities = 276/391 (70%), Positives = 323/391 (82%), Gaps = 3/391 (0%) |
|
| Structural Features: | pfam03573, OprD, outer membrane porin, OprD family.
CD-Length = 449 residues, 96.9% aligned
Score = 253 bits (648), Expect = 2e-68
Identities = 149/436 (34%), Positives = 208/436 (47%), Gaps = 47/436 (10%) |
|
|
|
|
|
| Homologs By Global Alignment |
| Gene ID:1229 |
Identity: |
0% |
10% |
20% |
30% |
40% |
50% |
60% |
70% |
80% |
90% |
100% |
|
|---|
| HomologID |
Accession |
Description |
Length |
PctIdentity |
PctSimilarity |
Gaps |
Score |
|---|
| 2234 |
PA0240_tr |
translation of PA0240 |
421 |
99.52 |
99.52 |
0 |
2222.0 |
| 73561 |
gb|AAG03629.1|AE004462_2 |
(AE004462) probable porin [Pseudomonas aeruginosa PAO1] |
421 |
99.52 |
99.52 |
0 |
2222.0 |
| 73562 |
gb|AAD39555.1| |
(AF031417) PhaK-like protein [Pseudomonas putida] |
424 |
69.81 |
81.83 |
9 |
1545.0 |
| 73563 |
gb|AAN67006.1|AE016329_1 |
(AE016779) BenF-like porin [Pseudomonas putida KT2440] |
424 |
69.57 |
81.83 |
9 |
1542.0 |
| 73565 |
gb|AAQ92184.1| |
(AY349612) PhaK-like protein [Pseudomonas fluorescens] |
426 |
67.13 |
78.40 |
12 |
1481.0 |
|
| GC ORFID: 44231 | How Found: BLASTX |
| GC_TrimmedSeqID: 69 | Blast Result ID |
| Subject Sequence Name: t_PA0240 | Glimmer Score: |
| Start: 6062980 | Stop: 6061715 |
| Length: 1266 | |
| Start Codon: ATG | Truncated Start: |
| Stop Codon: TGA | Truncated Stop: |
| Homolog: t_PA0240 translation of PA0240 | Homolog Bit Score 860.0 |
| Other Homologs: psyr_15may02_Scaffold1_revised_gene342 (ORF: BLASTX 580.0), psyr_15may02_Scaffold3_revised_gene5124 (ORF: BLASTX 565.0), t_PA4898 (ORF: BLASTX 440.0), t_PA2213 (ORF: BLASTX 410.0), t_PA0755 (ORF: BLASTX 331.0), t_PA3038 (ORF: BLASTX 312.0), t_PA4137 (ORF: BLASTX 302.0), t_PA2113 (ORF: BLASTX 296.0), t_PA1025 (ORF: BLASTX 296.0), t_PA3588 (ORF: BLASTX 293.0), t_PA4179 (ORF: BLASTX 290.0), psyr_15may02_Scaffold1_revised_gene1320 (ORF: BLASTX 278.0), t_PA0240 (ORF: GLIMMER) |
| GC ORF Sequence |
ATGTCGCACCCGAGCCACGTCCGCGCGCTCTGCGCTTCCCTCTGCCTGGGCGCCGGCCTGCCAGTCCATGCC
GGGCACGTCCATGCCGGGCAAGGTTTCCTCGAAGATGCCAAGGCCAGCCTCACCGCGCGCAATTTCCACCTG
CACCGCAACTTCGTCGGCGATGCCAGCCAGGGCAAGGCCGAGGAGTGGACGCAGAGCTTCATCCTCGACGCC
CGCTCCGGCTTCACCCAGGGCAGCGTCGGCTTCGGCCTCGATGTCCTCGGTCTCTACAGCCTGAAGCTGGAT
GGCGGCAAGGGAACGGCCGGCACCCAACTGCTGCCAACCCACGACGACGGCCGCCCGGCCGACGACTTCGGA
CGCCTGGCGGTGGCCGGCAAGCTGCGCGTGTCGAACAGCGAACTGAAGATCGGCGAATGGATGCCGGTGCTG
CCGATCCTCCGCTCCGACGATGGCCGCTCGCTGCCGCAGACCTTCCGCGGCGGCCAGCTCAGCGCCAACGAG
ATCGCCGGCCTGACCCTCTACGCCGGCCAGTTCCGCGGCAACAGCCCGCGCAACGACGCAAGCATGCAGGAC
ATGTCGCTGTTCGGCCGGCCAACCGCCACTTCCGACCGCTTCGATTTCGCCGGCGGCGAATATCGCTTCAAT
GGCGAACGCAGCCTGCTCGGCCTGTGGAACGCCGAGCTGAAGGACATCTACCGCCAACAGTACCTGCAACTG
CAACACAGCCAGCCGCTCGGCGACTGGCTGCTGGGCGCCAACCTCGGCGGCTTCCGTGGCCGCGACGCTGGC
AGCGCGCGCGCCGGCAAGCTCGATAACCGCACCGTTTCCGCGCTGTTCTCCGCGCGCTACGGGCTGCATACC
CTCTATCTCGGCCTGCAGAAGGTCAGTGGCGACGATGGCTGGATGCGGGTCAACGGCACCAGCGGCGGAACC
CTGGCCAACGACAGCTACAACGCCAGCTACGACAATCCCGGCGAGCGCTCCTGGCAACTGCGCTACGACTTC
GACTTCGTTGGCCTCGGCCTACCCGGCCTGACCTTCATGACCCGCTACCTGCATGGCGACCATGTGCGCCTG
GCCGGAGTCACCGACGACGGCAGCGAATGGGGACGCGAGAGCGAGCTGGGCTACACGCTGCAGAGCGGCGCC
TTCAAGCGTCTCAACGTCCGCTGGCGCAACTCCAGCCAGCGCCGCGACTGGGGCAGCAATACCCGCTTCGAC
GAGAACCGGCTGATCGTCAGCTATCCGCTGAGCCTGTTGTGA |
|